Supercomputers Target Coronavirus
Scientists at Oak Ridge and in China are putting their compute resources to work against the pandemic.

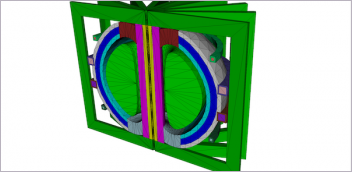
The compound in this image (in gray) was calculated to bind to the SARS-CoV-2 spike protein (cyan) to prevent it from docking to the human ACE2 receptor (in purple). Image courtesy of Micholas Smith/ORNL, U.S. Department of Energy.
Latest News
March 17, 2020
As governments around the world scramble to contain the coronavirus pandemic by shutting down schools, canceling public events, limiting travel, and ramping up testing efforts, powerful technology resources are also being deployed.
At Oak Ridge National Laboratory (ORNL) in Tennessee, researchers used the Summit supercomputer to identify 77 small-molecule drug compounds that could potentially bind to the glycosylated spike (S) protein, through which the COVID-19 virus accesses host cells. One or more of those compounds could potentially disarm the virus and point the way to future treatments.
“We were able to design a thorough computational model based on information that has only recently been published in the literature on this virus,” said team member and UT/ORNL CMB postdoctoral researcher Micholas Smith. Smith was referring to a previous study published in Science China Life Sciences that modeled the spike protein for risk of human transmission. When Chinese researchers sequenced the virus, they found that it infects the body via a mechanism similar to that of the SARS virus.
The power of the Summit supercomputer made it possible to rapidly evaluate a large number of potential compounds.
“In any situation where time is a critical factor—for example, a health issue or a company trying to get a new product out to market—supercomputers can play a critical role through digital simulation. It would be prohibitively time-consuming for researchers to grow virus cultures and test them with the near infinite amount of molecular compounds that exist in the physical world,” Dave Turek, IBM’s Vice President, Exascale Systems, told Nextgov in a recent interview. “So supercomputers like Summit can speed up that process by digitally simulating the effect to narrow down the range of possibilities researchers can look into.”
According to the South China Morning Post, researchers in China are also leveraging the Tianhe-1 supercomputer and artificial intelligence (AI) to diagnose COVID-19 patients using chest scans. The National Supercomputer Centre in Tianjin claims the system was able to provide a diagnosis wihtin 10 seconds based on CT scan images. The system can direct doctors to other areas of concern in a patient's lungs, provides an estimate of the likelihood of the person having COVID-19, and even provides treatment suggestions.
The idea of using CT scans for diagnosis first emerged in the Wuhan province, proposed as a faster way to get a diagnosis compared to genetic testing methods that could take several days. However, the U.S. Centers for Disease Control does not recommend a scan-based method, because protocols in the U.S. would require cleaning the scanning equipment after each use. In addition, the influx of suspected coronavirus patients into the hospital for scanning could potentially increase infection rates.
Modeling Coronavirus
At ORNL Smith built a model of the coronavirus spike protein based on the earlier studies of the structure. The Summit computer used NVIDIA V100 GPUs and GROMACS, a GPU-accelerated simulation package for biomolecular systems, to run the team’s experiments on more than 8,000 compounds. The results were published last month on ChemRxiv, a preprint server for chemistry research.
Smith was granted computational time on Summit through a Director’s Discretionary allocation, and he used a chemical simulations code to perform molecular dynamics simulations. Those simulations analyze the movements of atoms and particles in the protein. The team simulated different compounds docking to the S-protein spike of the coronavirus to determine if they could potentially prevent the spike from sticking to human cells.
“Using Summit, we ranked these compounds based on a set of criteria related to how likely they were to bind to the S-protein spike,” Smith said.
The team identified 77 small-molecule compounds, including medications and natural compounds, that could be candidates for further experimental testing. In the simulations, the compounds bind to regions of the spike that are important for entry into the human cell.
Researchers at the University of Texas at Austin have since released a more accurate model of the S protein, and the ORNL researchers plan to use that model to re-run the study.
“Summit was needed to rapidly get the simulation results we needed. It took us a day or two whereas it would have taken months on a normal computer,” said Jeremy C. Smith, Governor’s Chair at the University of Tennessee and director of the UT/ORNL Center for Molecular Biophysics. “Our results don’t mean that we have found a cure or treatment for the Wuhan coronavirus. We are very hopeful, though, that our computational findings will both inform future studies and provide a framework that experimentalists will use to further investigate these compounds. Only then will we know whether any of them exhibit the characteristics needed to mitigate this virus.”
IBM, which built the Summit, is also providing other resources to help researchers address the pandemic. According to IBM, the company’s Watson Health unit is working directly with health organizations around the world to better understand the nature of COVID-19. The IBM Clinical Development system has been made available without charge to national health agencies to accelerate clinical trials by providing data and analysis from web-enabled devices. In addition, IBM’s Operational Risk Insight tool has been made available to non-profit organizations to help analyze data on the spread of the virus.
You can read more about the ORNL and Department of Energy activities related to coronavirus at the Coronavirus Hub. ORNL has also posted a video about its research here.
Subscribe to our FREE magazine, FREE email newsletters or both!
Latest News
About the Author
DE’s editors contribute news and new product announcements to Digital Engineering.
Press releases may be sent to them via DE-Editors@digitaleng.news.